What is pathway analysis gene?
What is pathway analysis gene?
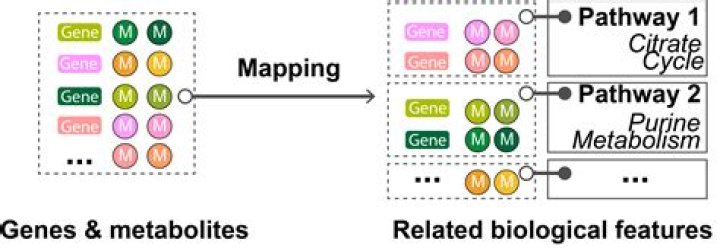
Pathway enrichment analysis helps researchers gain mechanistic insight into gene lists generated from genome-scale (omics) experiments. This method identifies biological pathways that are enriched in a gene list more than would be expected by chance.
What is the use of pathway analysis?
Pathway analysis is a set of widely used tools for research in life sciences intended to give meaning to high-throughput biological data. The methodology of these tools settles in the gathering and usage of knowledge that comprise biomolecular functioning, coupled with statistical testing and other algorithms.
How do you find genes in a pathway?
Use the advanced gene query by selecting Genes under the Search menu. You may enter either the name of the pathway (“Alzheimer’s disease”) or the KEGG or REACTOME ID of the term (e.g., “KEGG:05010”).
How do you use GSEA?
The basic steps for running an analysis in GSEA are as follows:
- Prepare your data files: ▪ Expression dataset file (res, gct, pcl, or txt) ▪ Phenotype labels file (cls)
- Load your data files into GSEA. See Loading Data.
- Set the analysis parameters and run the analysis. See Running Analyses.
- View the analysis results.
How do you use a G Profiler?
g:Profiler Tutorial
- Step 1: Generate g:Profiler output files. Go to g:Profiler website. Select and copy all genes in the tutorial file 12hr_topgenes.
- Step 2: Generate Enrichment Map with g:Profiler Output. g:Profiler output files from Step 1: gProfiler_EM.zip. Open Cytoscape.
- Step 3: Examining Results. Legend:
What is the difference between GO and KEGG?
GO stands for Gene Ontology and as the name suggests, it annotates genes using an ontology. KEGG, Panther and other “pathway” databases group genes into “pathways” which are basically lists of genes participating in the same biological process.
Which bioinformatics software can you use for pathway analysis?
Reactome is a free, open-source, curated and peer-reviewed pathway database. Our goal is to provide intuitive bioinformatics tools for the visualization, interpretation and analysis of pathway knowledge to support basic research, genome analysis, modeling, systems biology and education.
What is the difference between KEGG and Reactome?
Both KEGG and Reactome covers same number of genes ( example for human ~7000). The difference is KEGG has more broad term and Reactome has similar terms but as multiple detailed entries (splited terms for same entry from KEGG) .
What is Pathway database in bioinformatics?
A pathway can be viewed as a set of interconnected processes, while a process can be viewed as being made up of molecular entities. More generally, an entire pathways database can be viewed as a single large graph of interconnected reactions, in which certain subgraphs are identified as specific pathways.
What is pathway analysis in bioinformatics?
In bioinformatics, methods of pathway analysis might be used to identify key genes/ proteins within a previously known pathway in relation to a particular experiment / pathological condition or building a pathway de novo from proteins that have been identified as key affected elements. …
How do you analyze data on GSEA?
How do you use a KEGG Mapper?
You can use the KEGG database directly @ Click on the KEGG mapping displayed on the left side, then click on the search pathway, and paste the gene ID in the displayed box. finally, execute to get the results of your analysis. It is user friendly and works perfectly.


